On March 28–29, 2026, the 4th International Conference on Single-Cell and Spatial Omics (TICSSO-4), themed “Atlas AI Acceleration,” was successfully held in Beijing. Leading scholars, industry professionals, and young scientists from around the world gathered to discuss technological breakthroughs and industrial applications in single-cell and spatial omics. Professor Sunney Xie, founder of Cygnus Biosciences, and Professor Yanyi Huang were invited to attend the conference and shared Cygnus Biosciences latest advancements in single-cell sequencing.
Professor Sunney Xie delivered the opening keynote speech titled “What Can FOODIE Do for Biology and Medicine,” while Professor Yanyi Huang presented a systematic overview of Cygnus Biosciences proprietary technology framework—built from first-principles—under the theme “Subcellular-Resolution Transcript Spatial Localization Analysis.”

Figure 1: Professor Sunney Xie, Director of the Changping Laboratory, delivering a keynote speech

Figure 2: Professor Yanyi Huang from Peking University delivering a keynote speech
Cygnus Biosciences’s technological innovation begins with breakthroughs in foundational technologies. Building upon Fluorogenic sequencing chemistry and ECC error-correction coding, Cygnus Biosciences has established two core pillars: LumoSeq sequencing chemistry and AttoSeal microfluidic manipulation technology. Together, these support the unique advantages of the GS200 and GP1000 gene sequencers in the fields of single-cell transcriptomics and spatial transcriptomics.
The LumoSeq sequencing chemistry employs a novel nucleotide molecule. Leveraging its unique ability to switch between fluorescent on and off states, it enables the detection of all four nucleotide bases using only a single-color fluorescence, without leaving molecular scars on the DNA. This feature not only simplifies instrument design but also eliminates cumbersome operational steps such as adding balance libraries, making it suitable for a wider range of applications. AttoSeal microfluidic control technology enables precise fluid management of micro-reactors at the attoliter (10⁻¹⁸ L) scale, allowing sequencing biochemical reactions to occur in parallel and synchronously across nearly 10 billion micro-reactors. This significantly reduces the consumption of sequencing reagents, thereby dramatically lowering sequencing costs.
Taking the GP1000 gene sequencer as an example, it has demonstrated exceptional performance and capabilities in the field of single-cell sequencing. Recent experimental data has further robustly validated the GP1000 platform’s outstanding performance in single-cell transcriptomics.
Researchers utilized the GP1000 platform to perform library sequencing based on 10x Genomics’ 3’ scRNA-seq technology, conducting a comparative analysis of Resident Memory T cells (Trm) generated by acute infections and Transitory Exhausted T cells (Ttrans) associated with chronic infections.
Test data show that after processing the raw data generated by the GP1000 and performing dimensionality reduction analysis using the UMAP algorithm (Figure 3), the samples exhibited distinct clustering patterns.
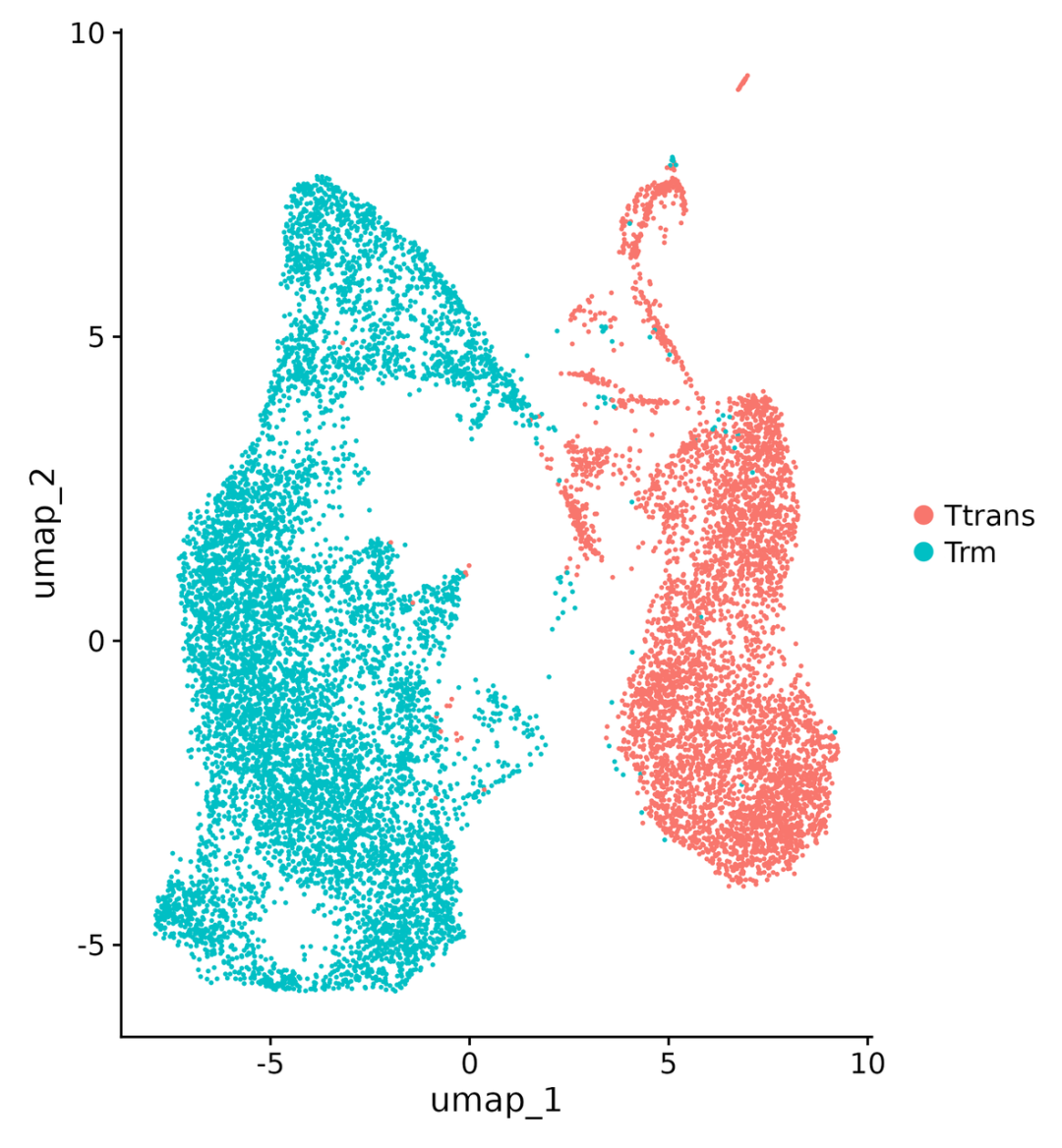
Figure 3: UMAP dimensionality reduction of single-cell transcriptomics and cell types
The light green cluster in the figure represents Trm derived from acute infections, while the red cluster represents Ttrans from chronic infections. The two cell types form distinct morphological distinctions in the UMAP space, indicating significant differences in their gene expression. This demonstrates that the transcriptional features captured by the GP1000 are sufficient to support high-resolution cell subtype clustering.
The distribution of marker gene expression shown in Figure 4 demonstrates that the high-quality data generated by GP1000 not only confirms classic cellular identity markers but also more sensitively captures the transcriptional plasticity of T cells during differentiation:
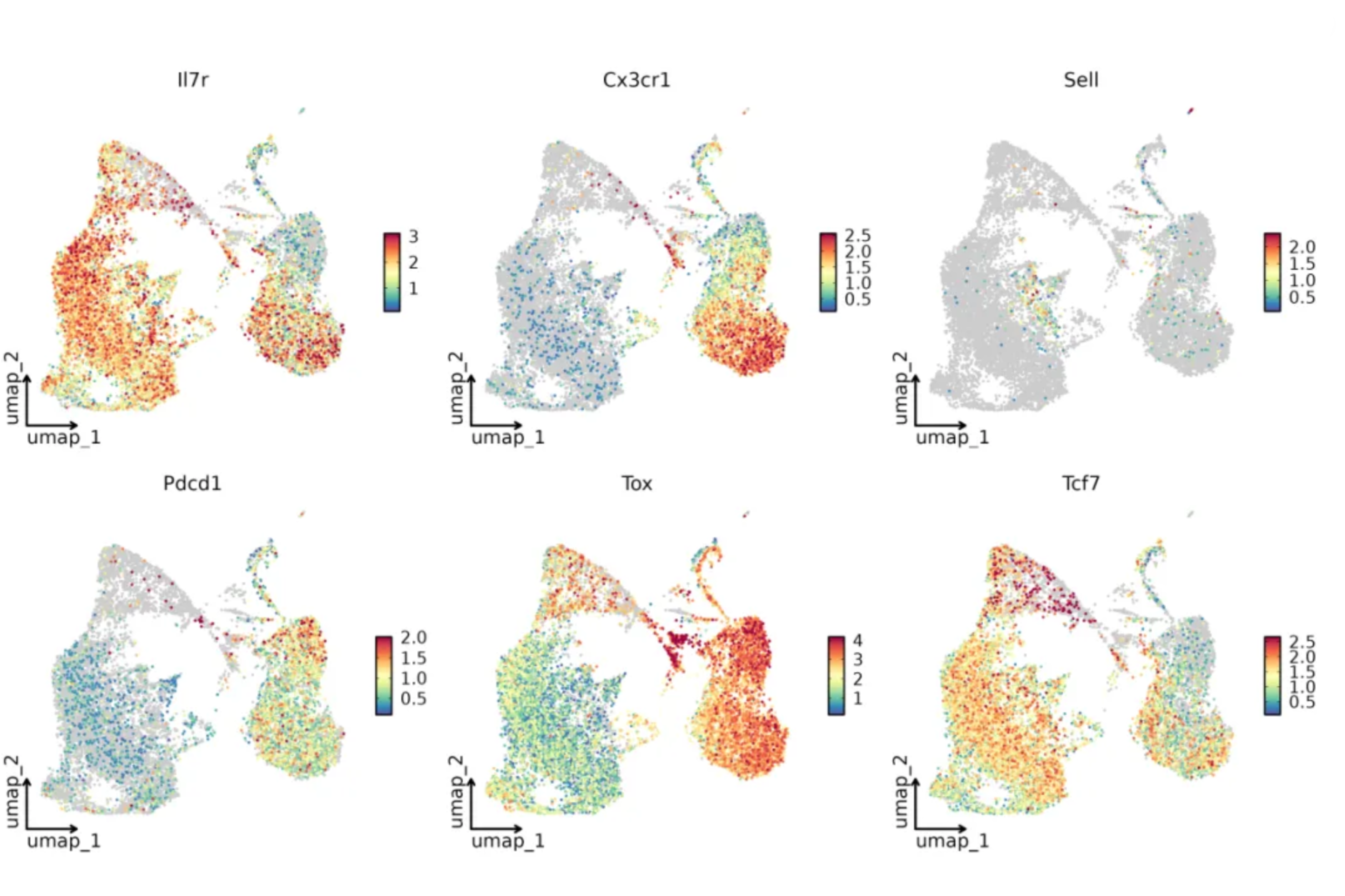
Figure 4: Distribution of marker gene expression
Leveraging the high-resolution capabilities of single-cell sequencing, the data precisely reveal the antagonistic characteristics of Tcf7 and Tox within the Ttrans population, as well as the transcriptional gradient leading from precursor-exhausted T cells (Tpex) to exhausted T cells (Tex). Concurrently, the heterogeneous co-expression of Il7r and Cx3cr1 further reflects the functional subpopulation heterogeneity within this transitional state. The consistently low expression of Sell in both populations validates, at the transcriptional level, its characteristics as a transitional state between tissue residency and activation.
These detailed transcriptomic differential analyses demonstrate that the GP1000 gene sequencer possesses the ability to capture subtle fluctuations within cell populations—for Ttrans cells in a dynamic transitional phase at the transcriptional level, the GP1000 provides sufficient digital evidence to support high-resolution subpopulation differentiation. These results have also received high acclaim from industry experts.
The presentations by Professor Sunney Xie and Professor Yanyi Huang sparked an enthusiastic response from attending scholars at the conference. Several authoritative experts highly praised Cygnus Biosciences’s end-to-end, proprietary technology roadmap, which spans from fluorogenic sequencing chemistry, ECC error-correction coding, and LumoSeq sequencing chemistry to AttoSeal microfluidic manipulation. Cygnus Biosciences’s independently developed gene sequencing technology has achieved major breakthroughs in sensitivity, accuracy, and cost control. Its independently controlled sequencing platform holds critical value in advancing the progress of life science research.





