As biomedical research enters the single-cell era, high-throughput sequencing depth and high-quality data output have become the cornerstone for dissecting complex cellular landscapes. Under its OpenSeq Initiative, Cygnus Biosciences (Beijing) Co., Ltd. aims to support global researchers in exploring cutting-edge life science fields using the GP1000 (P1000) Sequencing System by providing a total of 1,000 terabytes of free sequencing data.
Recently, researchers employed the Cygnus Bio GP1000 platform to sequence libraries prepared with 10x Genomics 3' scRNA-seq technology, where the GP1000 delivered outstanding sequencing performance in complex immune cell populations.
Research Background: Comparative Analysis of T-Cell Subsets
The test samples focused on two critical T-cell states: resident memory T cells (Trm) generated during acute infection and transitory exhausted T cells (Ttrans) observed in chronic infection.
In immunological research, T-cell states determine the body’s efficacy against pathogens or tumors. Trm cells reside long-term in tissues and serve as the first line of defense against reinfection, whereas Ttrans cells represent a key transitional population as T cells evolve from functional effector states to terminal exhaustion under chronic antigen stimulation. Deciphering the transcriptomic differences between these two cell types carries profound biological significance for understanding the immunopathology of chronic viral infections and developing targeted immunotherapies [1].
Sequencing Quality: Dual Assurance of Depth and Precision
As illustrated in Figure 1, the GP1000 sequencer demonstrated exceptional data quality and output stability:
These metrics confirm that the GP1000 is fully compatible with mainstream single-cell library preparation platforms such as 10x Genomics, delivering high-fidelity raw data for downstream analysis.
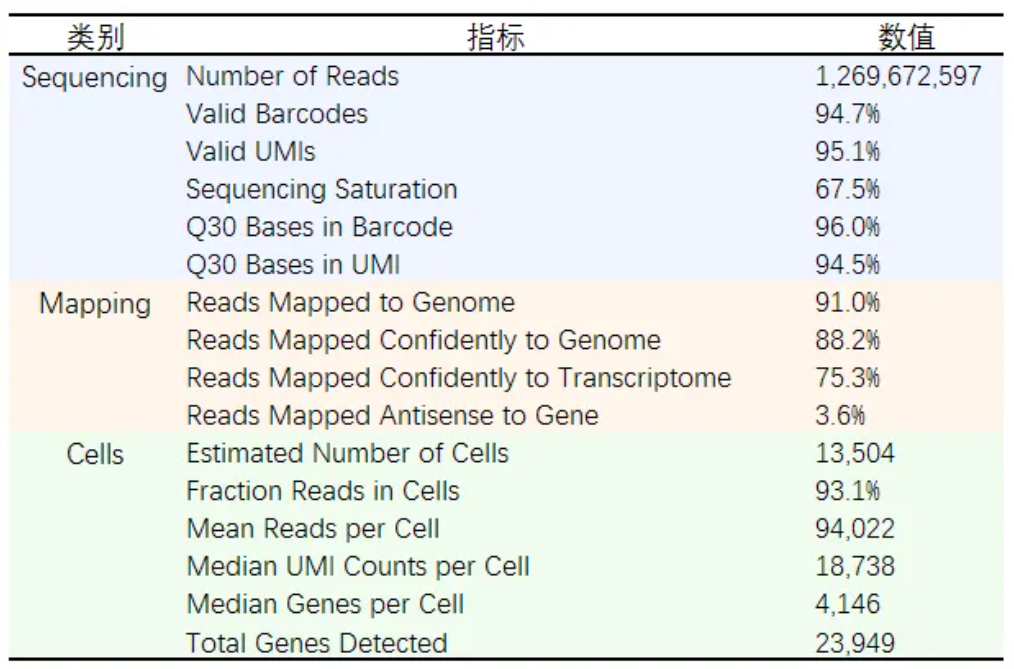
Table 1: Quality Control Metrics from CellRanger Software
Cell Clustering: Clear Profiling of the Immune Landscape
After processing raw data generated by the GP1000, dimensionality reduction analysis using the UMAP algorithm (Figure 1) revealed distinct population clustering. The turquoise cluster represents Trm cells derived from acute infection, while the red cluster denotes Ttrans cells in chronic infection. The two cell types showed clear spatial separation in the UMAP plot, indicating significant disparities in gene expression and proving that the transcriptomic signatures captured by the GP1000 support high-resolution cell subset classification.
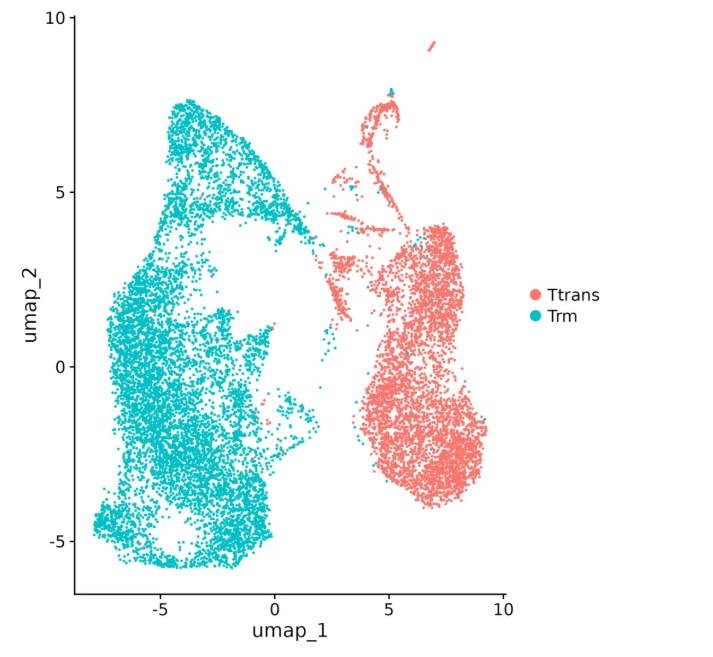
Figure 1: UMAP Dimensionality Reduction and Cell Typing of Single-Cell Transcriptomes
Marker Gene Validation: Accurate Identification of Functional States
Supported by the marker gene expression profiles in Figure 2, the high-quality data produced by the GP1000 not only validated canonical cell identity features but also sensitively detected transcriptional plasticity during T-cell differentiation:
Such refined transcriptomic differential analysis demonstrates that the GP1000 sequencing system can capture subtle fluctuations within cell populations. For Ttrans cells at dynamically transitional transcriptional stages, the GP1000 provides robust digital evidence supporting high-resolution subset delineation.
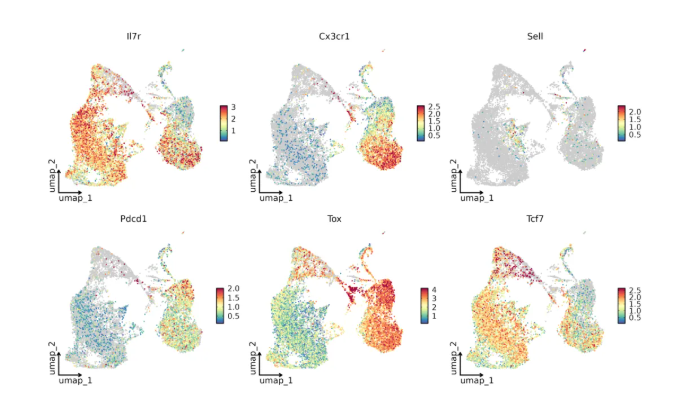
Figure 2: Marker Gene Expression Distribution
Conclusion: Join the OpenSeq Initiative to Explore the Future Together
This real-world test showcases the technological advantages of the Cygnus Bio GP1000 in complex single-cell transcriptome research. From rigorous quality control metrics to sensitive detection of intermediate cell characteristics, the GP1000 serves as a reliable tool for immunological research.
The OpenSeq Initiative is now accepting applications! We look forward to partnering with more research colleagues to unlock the mysteries of the single-cell world. Whether your work focuses on developmental biology, tumor immunity, or neuroscience, Cygnus Bio will provide powerful high-throughput sequencing support.
Contact us directly.Email: OpenSeq@cygnusbio.com
References





