Recently, a research achievement titled "A fuzzy sequencer for rapid DNA fragment counting and genotyping" was published in Nature Biomedical Engineering by a team led by Professor Huang Yanyi from Peking University, a team led by Professor Wang Jianbin from Tsinghua University, and Cygnus Bio. The study innovatively proposed a fuzzy sequencing technology, which can accurately align the sequenced DNA fragments to the reference genome without obtaining the exact four-base DNA sequence. This innovative breakthrough enables faster resequencing analysis results, greatly improving the efficiency of DNA analysis and reducing sequencing costs.
Fuzzy sequencing technology includes two types: BitSeq and SuperBitSeq. Compared with traditional sequencing methods, both can provide higher information entropy and read length under the same number of sequencing cycles. Specifically, BitSeq provides fuzzy sequences composed of M (A or C) and K (G or T). The information entropy generated per cycle is equivalent to that of reversible terminator sequencing (both 2 bit/cycle), but the read length is twice that of the latter (2 bp/cycle). SuperBitSeq generates sequences composed of four bases but with uncertain local base ordering. Its read length per cycle is equivalent to that of BitSeq, and the information entropy is increased to 3.37 bit/cycle. In simulated alignment experiments conducted on human and Arabidopsis thaliana genomes, the results showed that to achieve the same unique alignment rate, both BitSeq and SuperBitSeq required significantly fewer sequencing cycles than reversible terminator sequencing.
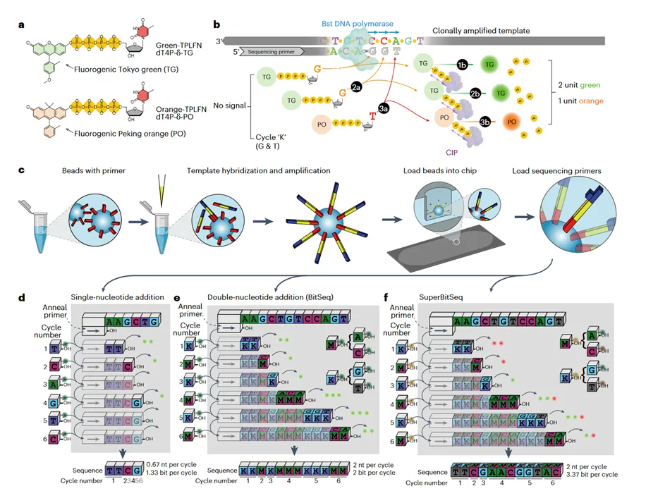
In the field of biological and medical research, high-throughput sequencing technology has become an indispensable tool, with its application scope continuously expanding. Among numerous application scenarios, resequencing occupies an absolute dominant position. Based on the known reference genome, resequencing accurately aligns the sequenced sequences to the reference genome, and then infers the sample composition based on a large number of sequence alignment results. Typical resequencing includes copy number variation (CNV) detection, non-invasive prenatal testing (NIPT), RNA sequencing (RNA-seq), metagenomic sequencing (mNGS), etc.
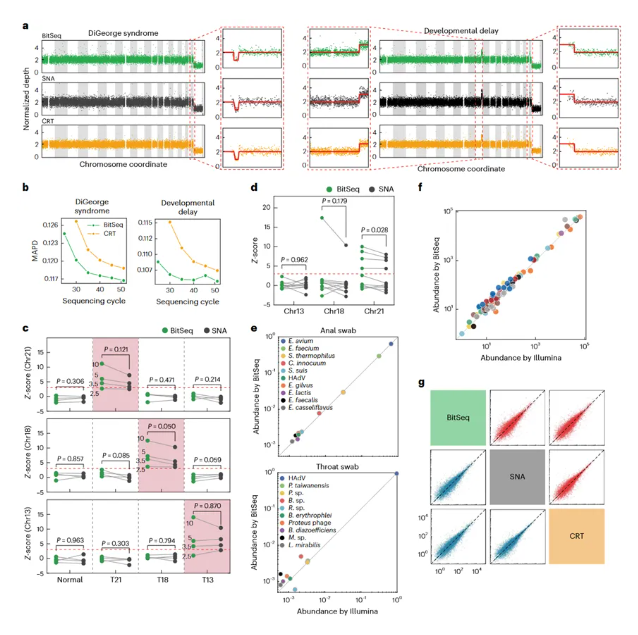
Among the many applications of resequencing, counting-based applications are a key category, whose core is to count the number of different types of nucleic acids. The characteristics of BitSeq fuzzy sequencing are highly consistent with the needs of counting-based applications. The research team conducted tests on counting-based application scenarios such as CNV, NIPT, RNA-seq, and mNGS. The results showed that the results obtained by BitSeq fuzzy sequencing technology were highly consistent with those of reversible terminator sequencing and semiconductor sequencing, demonstrating its great application potential in this field.
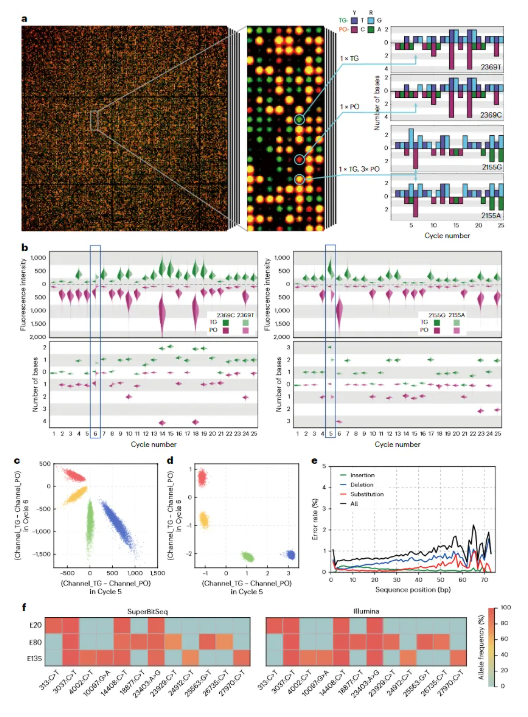
On the basis of BitSeq, SuperBitSeq shows more powerful application potential. It is not only applicable to counting-based applications such as CNV, NIPT, RNA-seq, and mNGS, but also breaks through the ability to detect single nucleotide variations (SNVs). The research team further realized high-throughput SuperBitSeq, successfully distinguishing the two mutant sequences G719S and T790M of the EGFR gene and their corresponding wild-type sequences. In the detection of COVID-19 SNVs, its results were also highly consistent with traditional methods.
As a domestic original research enterprise in gene sequencing technology, Cygnus Bio has two fully independent intellectual property sequencing system product lines: Product Line S and Product Line P. Product Line S sequencing system includes two models, GS100 and GS200, which have successfully achieved full compatibility with BitSeq fuzzy sequencing technology. Relying on the advantages of fuzzy sequencing technology, Product Line S sequencing system will inject new momentum into the fields of biological and medical research.





